
Code
SynEM is an algorithm for the detection of synapses from 3d electron microscopy images for connectomics. All SynEM code is available via a repository on gitlab released under the MIT license. The code base on gitlab may contain improvements and bugfixes to the SynEM supplementary code supplied in the paper at eLife. We therefore recommend using the code version on github.
More ›
Data/SynEM challenge
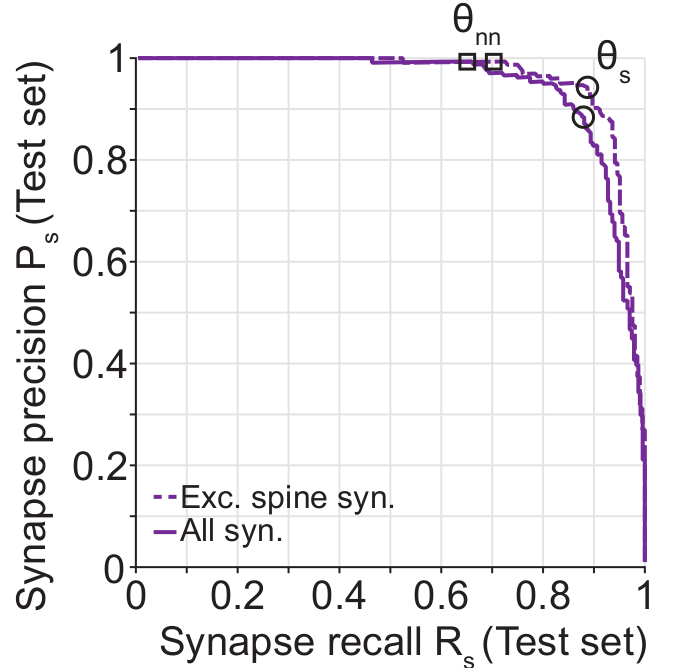
The data that was used to train and test SynEM as well as the modified segmentation for the Kasthur et al. (2015) dataset is available in a WebDAV repository. To compare algorithms on the SynEM challenge, the provided data with the routines in the SynEM code are sufficient (see code file +SynEM/+Eval/interfaceRP.m).
More ›
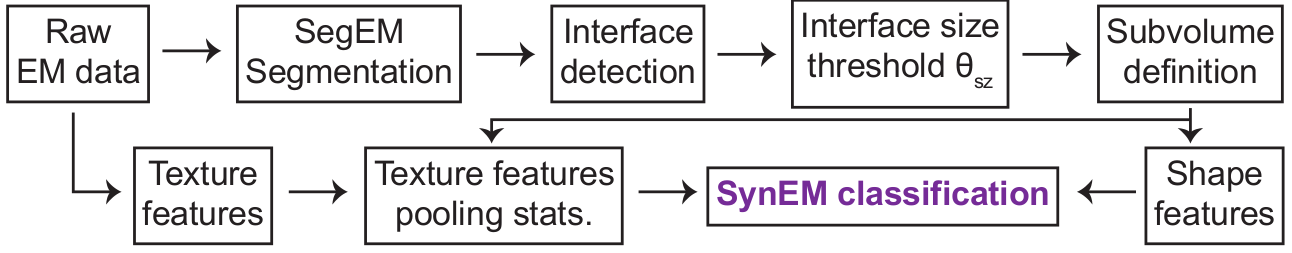
Integrated SegEM - SynEM - webknossos workflow
SynEM is an automated method for synapse detection in 3D EM data. For the reconstruction of neurites (the "wires") in such data, we developed SegEM and webKnossos
More ›